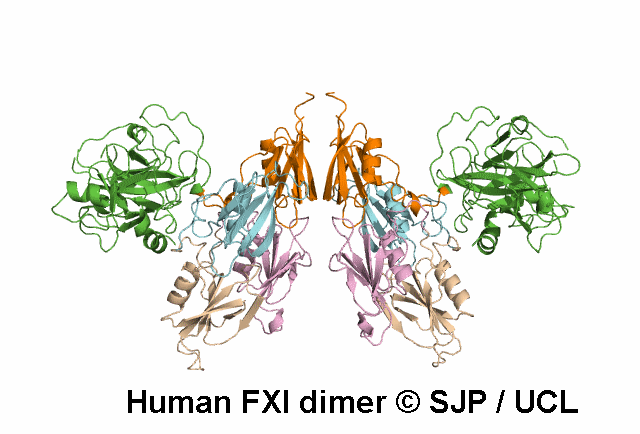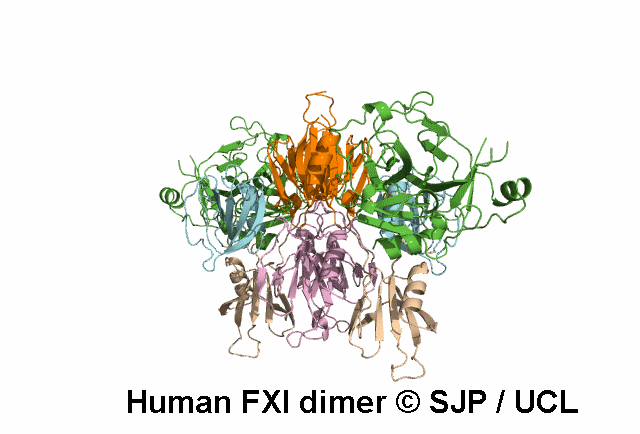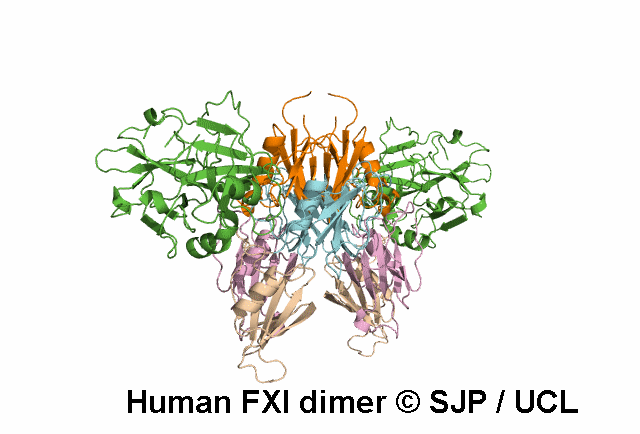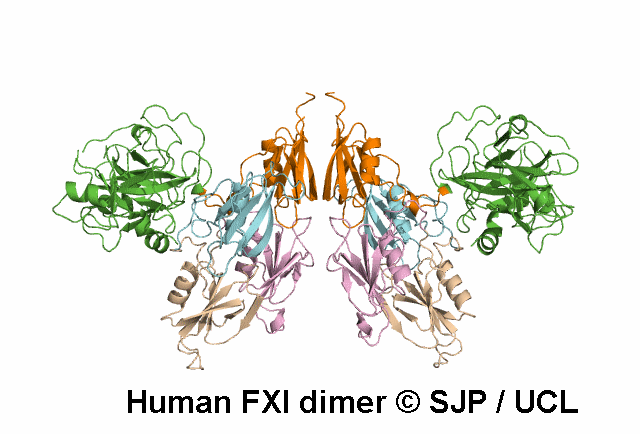In Depth Variant Analysis: c.1336A>G (p.Ser446Gly)
Terms with a '*' next to them are explained on the Help Page .
c.1336A>G
p.Ser446Gly (Legacy AA No. 428)
Variant Type:
Point
Domain:
Serine Protease
Codon Change:
A>G
Variant Effect:
Missense
No. of Patients Reported:
0
Phenotype:
None
Allele Count *:
2
Allele Number *:
251392
Allele Frequency *:
0.000008
Variant Comments & Reference:
One small hydrophilic amino acid (Ser) is replaced by another small, hydrophilic amino acid (Gly). Ser446 is conserved in bovine FXI and human prekallikrein, but it is not conserved in rabbit or murine FXI. Likely to be a non-causative polymorphism. Duncan et al 2008Residue Information:
| Name | Type | Cyclic | Size | Position | Hydrophobicity | Charge | |
|---|---|---|---|---|---|---|---|
| |
|
|
|
|
|
|
|
| |
|
|
|
|
|
|
|
Substitution Analysis:
Structural Implications:
Ser446 is a buried residue (surface accessibility value = 0 ).
Ser446 is in a region of secondary structure within the FXI domains.
The DSSP assignment for this residue is ... E.
Ser446 is in a region of secondary structure within the FXI domains.
The DSSP assignment for this residue is ... E.
Hint: Left click mouse | To rotate the structure
Hint: Rotate mousewheel | To zoom in/out the structure
Hint: Right click mouse | to use applet control options
|
| Spacefill | |
| Cartoon | |
| Wireframe | |
| Trace | |
| Backbone | |
| Spin | |
| Background | |
| Disulphides | |
| Domains | |
| Alternative Colouring | |
| Labels |
Right Click on the molecule's screen for more options.
NOTE: If no Java applet appears or a Java error message is shown, please click the following LINK.